Research
The patterns of muscle activation that drive mammalian movement reflect an interplay between networks of neurons in the spinal cord and neurons in an array of brain regions that comprise the motor system. However, current models of motor system operation say little about this interplay on the short timescales over which neurons communicate and movement is generated. Our research combines novel genetically-mediated approaches for measuring and perturbing activity in neuronal subpopulations with quantitative and model-based analyses to understand this interplay. Here are some examples of our current interests:
Skilled movement generation in real time – Motor skills are movements that animals improve with repeated practice. The capacity to improve motor behaviors is vital to the fitness these behaviors confer, and so to the evolutionary success of many mammalian species. Many motor system regions appear critical for driving motor skills, but it is unclear how these regions interact in real time to ensure such movements are properly orchestrated. Our approach is behavior-centric, exploring how neuronal populations within motor system regions conspire to execute a model skilled movement. We draw on our experience developing training paradigms in which mice learn skilled forelimb movements, which have opened up a wealth of experimental possibilities. We assess interactions between neuronal populations by combining rapid activity perturbations via optogenetic probes with large-scale electrical and optical activity measurement in one or more populations as movements are executed. Simultaneous forelimb electromyography (EMG) provides a high temporal resolution readout of motor system output.
Understanding the cortical orchestration of movement – Motor areas of the brain’s neocortex play a particular role in facilitating the remarkable complexity and agility of mammalian movement, but we understand little about how. In contrast to previous approaches that have assessed cortical function with respect to movement type, we focus on how motor cortical influence depends on the underlying muscle activation patterns themselves, which we quantify via EMG. Our approach relies on the temporal resolution of optogenetic inactivation methods, the development of freely-behaving paradigms in which mice express a broad range of natural movements, and the use of contemporary dimensionality reduction methods to classify muscle activation patterns. Our recent findings suggest that there are certain components of motor cortical output activity that directly drive muscles, while other components may serve other roles. Coupling activity measurement and perturbation here will allow us to further investigate this sort of functional discretization among motor cortical output components.
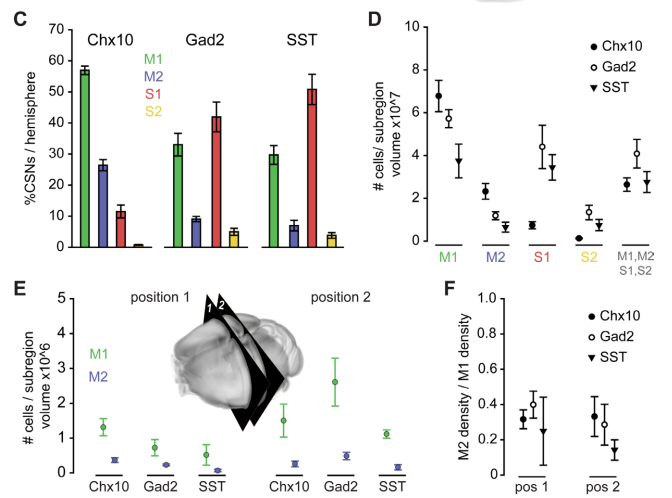
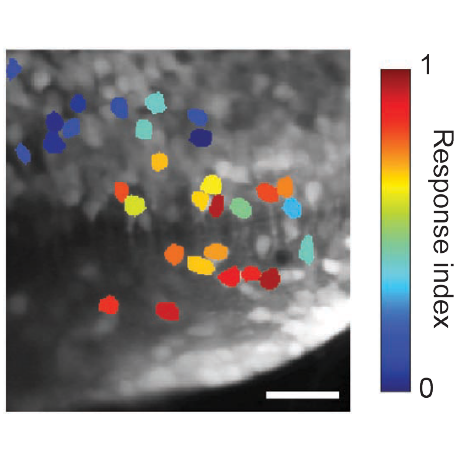
Identifying the functional units of corticospinal control – The resolution of subatomic particles or of DNA structure illustrate how mechanistic insight can rapidly mount after the appropriate functional units for explaining a given phenomenon are identified. However for many neural processes, the appropriate functional units for describing the underlying mechanism remain unclear. The continued development of viral tracing strategies and cellular-resolution genomic methods for delineating neuronal subtypes now enable us to address this ambiguity in a new way. We aim to characterize the relationship between neuronal identity and function within the corticospinal population by combining two-photon imaging of neural activity in mice performing skilled forelimb movements together with genetic access to subtypes defined by transcriptomic state, projection patterns, and postsynaptic partner identity. Dimensionality reduction-based analyses will enable unbiased assessment of coactive functional groups among corticospinal neurons that can be registered with subtype identity. By resolving how corticospinal neuron features align with function, light will be shed on the appropriate functional elements with which to describe motor system operation.
People
Andrew Miri, Ph.D.
Assistant Professor
Michaël Elbaz, Ph.D.
Postdoctoral Fellow
PhD, Systems Neuroscience
Zahra Amer
David Xing, Ph.D.
Mark Agrios
Graduate Student, Neuroscience. Co-advised by Dr. Sara Solla, Department of Neuroscience
Diya Basrai
Graduate Student, Neuroscience
Wentao Qiu
Graduate Student, Neurobiology
Zach Fitzgerald
Graduate Student, Neurobiology
Adithi Adusumilli
Undergraduate, Neuroscience & Computer Science Major
ALUMNI
Abhishek Sarup, M.S.
Diagnostic Lab Head, Rush University Medical Center
Adam Forrest
PhD student. Biomedical Engineering, University of Pittsburgh
Akiko Saiki Ph.D.
MATLAB engineer., Research Coordinate Inc.
Amy Kristl
Associate Medical Writer, Citrus Health Group
Daniel Greenberg
Hannah Sroussi
MD Student at Kaiser Permanente School of Medicine
Jason Xu
Kole Butterer
Research Technician
Meg Young
PhD student at Sainsbury Wellcome Centre for Neural Circuits and Behaviour (UCL)
Natalie Koh
Postdoc, Khanna Lab, UC Berkeley
Polina Cherepanova
PhD student, Princeton University
Sajishnu Savya
Data Scientist, Abbott Labs
Sarah Hsu
PhD student, Princeton University
Zhengyu Ma Ph.D.
Assistant Professor, AI Department, Peng Cheng National Laboratories
PREPRINTS
Natalie Koh, Zhengyu Ma, Abhishek Sarup, Amy C. Kristl, Mark Agrios, Margaret Young, Andrew Miri.
Samaher Fageiry, Claire L. Warriner, Jackson Loper, Liam Pianski, Thomas Reardon, Thomas M. Jessell, Rui M. Costa, Andrew Miri.
David Xing, Joshua Glaser, Andrew Miri
Amy Kristl, Natalie Koh, Mark Agrios, Sajishnu Savya, Zhengyu Ma, Diya Basrai, Sarah Hsu, Andrew Miri
selected Publications
Hierarchy in influence but not firing patterns among forelimb motor cortices.
Akiko Saiki-Ishikawa, Mark Agrios, Sajishnu Savya, Adam Forrest, Hannah Sroussi, Sarah Hsu, Diya Basrai, Feihong Xu, Andrew Miri (2025) Hierarchy between forelimb premotor and primary motor cortices and its manifestation in their firing patterns eLife 13:RP103069.
Motor cortical influence relies on task-specific activity covariation
Warriner, C.L., Fageiry, S., Saxena, S., Costa, R.M. and Miri, A. (2022) "Motor cortical influence relies on task-specific activity covariation." Cell Reports 40(13):111427.
Towards Cell and Subtype Resolved Functional Organization: Mouse as a Model for the Cortical Control of Movement
Warriner, C.L., Fageiry, S.K., Carmona, L.M., Miri, A. (2020) “Towards Cell and Subtype Resolved Functional Organization: Mouse as a Model for the Cortical Control of Movement.” Neuroscience 450:151-160.
Behaviorally-selective engagement of short-latency effector pathways by motor cortex.
Miri, A., Warriner, C.L., Seely, J.S., Elsayed, G.F., Cunningham, J.P., Churchland, M.M., Jessell, T.M. (2017). Neuron 95(3):683-696.
Primacy of flexor locomotor pattern revealed by ancestral reversion of motor neuron identity.
Machado, T.A., Pnevmatikakis, E., Paninski, L., Jessell, T.M., Miri, A. (2015). Cell 162(2):338-50.
Edging toward entelechy in motor control.
Miri, A., Azim, E., Jessell, T.M. (2013). Neuron 80(3):827-34.
Spatial gradients and
multidimensional dynamics in a neural integrator circuit.
Miri, A., Daie, K., Arrenberg, A.B., Baier, H., Aksay, E., Tank, D.W. (2011). Nature Neuroscience 14(9):1150-11.
Regression-based identification of behavior-encoding neurons during large scale optical imaging of neural activity at cellular resolution.
Miri, A., Daie, K., Burdine, R.D., Aksay, E., Tank, D.W. (2011). Journal of Neurophysiology 105(2):964-980.
Contact
Andrew Miri, Ph.D.
Assistant Professor
Department of Neurobiology
Northwestern University
2205 Tech Drive
2-115 Pancoe Hall
Evanston, IL 60201
andrewmiri@northwestern.edu